Human DPPI (also known as cathepsin C) is a ubiquitously expressed lysosomal peptidase with important physiological roles in the immune system, particularly for the activation of effector serine peptidases (granzymes, etc.). Therefore, it is a target for treatment of autoimmune and inflammatory diseases. Structure-wise, DPPI is special among papain-like peptidases because it contains an additional exclusion domain that is non-covalently attached to the catalytic domain (see image below). The exclusion domain restricts access of substrates into the active site and thereby determines the dipeptidyl-peptidase activity of DPPI. It also enables its tetramerization which is commonly considered as a unique feature of DPPI, as opposed to the monomeric structures of other papain-like peptidases.
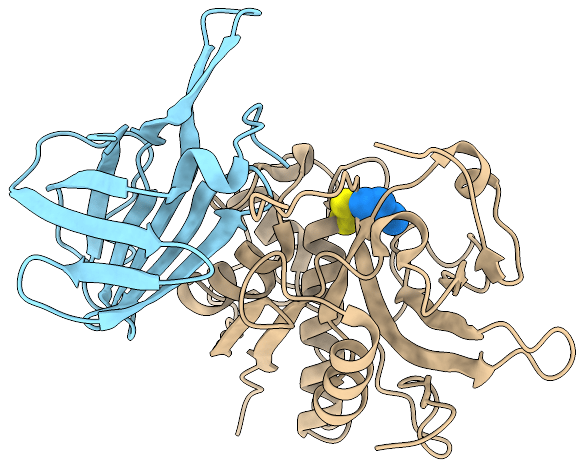
Crystal structure of a DPPI protomer. The peptidase domain is colored grey and the exclusion domain magenta (PDB ID 1K3B).
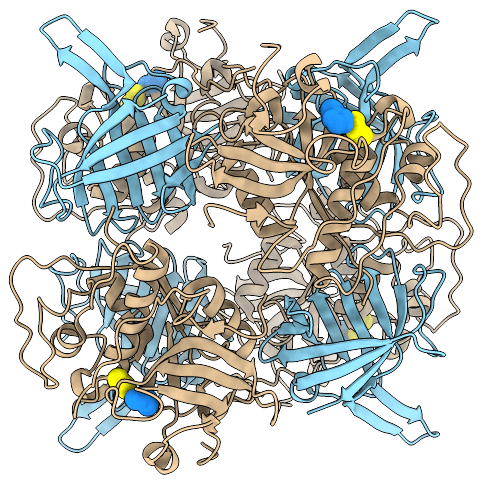
Crystal structure of the entire DPPI tetramer (PDB ID 1K3B).
We recently produced a monomeric form of DPPI which had activity nearly identical to the tetrameric form, demonstrating that tetramerization is not necessary for DPPI activity nor does it significantly affect its kinetic properties. Our goal is to understand the evolution of the DPPI tetramer and the implications of oligomerization for its functional properties. Based on currently available data we propose a stepwise model for the evolution and assembly of the DPPI tetramer:
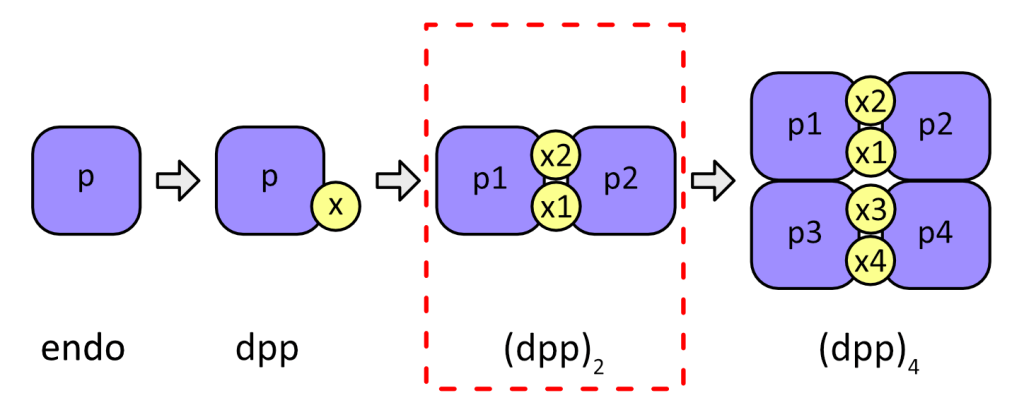
DPPI evolution began with the fusion of a regulatory (exclusion) domain with a papain-like endopeptidase (endo), generating a dipeptidyl-peptidase (dpp). A functional monomeric form reminiscent of this putative ancestor was described in Plasmodium. This initial monomeric form associated into dimers which are present in man at the zymogen level and finally into a tetrameric dimer of dimers. In mammals, proper N-glycosylation is necessary for maturation of DPPI in vivo, but mature DPPI no longer relies on N-linked oligosaccharides for stabilization and the oligosaccharide chains are for the most part not visible in crystal structures.
Our current goal is to verify the proposed model by phylogenetic and experimental analyses of DPPI homologs from divergent phylogenetic groups. Our recent phylogenetic analysis showed that DPPI is present in several eukaryote lineages, but is not ubiquitous in eukaryotes. In alveolates (e.g., Plasmodium) DPPIs are monomeric enzymes with significant insertions in comparison to their human counterpart and phylogenetic data suggest that oligomeric DPPIs emerged only in the Amorphea group, which contains animals, amoebae, etc. More details are found here.

